Code
# required packages
library(survival)
library(randomForestSRC)
library(midr)
library(ggplot2)
library(gridExtra)
library(patchwork)
data(flchain, package = "survival")We are thrilled to announce the release of {midr} 0.6.0. This update brings the core principles of the MID framework to a highly requested domain: Survival Analysis.
With this release, users can now decompose and interpret complex, “black-box” survival models—such as Random Survival Forests (RSF)—just as easily as standard regression and classification models.
Key new features in 0.6.0 include:
Native Support for Survival Models: The interpret() function now natively supports some survival models including Cox proportional hazard models ({survival}) and random survival forests ({randomForestSRC}, {ranger}).
Time-Dependent Sequence of MID Models: Because predicted survival probabilities change over time, {midr} now calculates and returns an array of time-varying MID surrogate models.
Here is a look at how {midr} 0.6.0 interprets a Random Survival Forest model using the flchain dataset.
Sample size: 7874
Number of deaths: 2169
Number of trees: 100
Forest terminal node size: 20
Average no. of terminal nodes: 245.64
No. of variables tried at each split: 3
Total no. of variables: 5
Resampling used to grow trees: swor
Resample size used to grow trees: 4976
Analysis: RSF
Family: surv
Splitting rule: logrank *random*
Number of random split points: 10
(OOB) CRPS: 463.86434184
(OOB) standardized CRPS: 0.09280999
(OOB) Requested performance error: 0.29755615
Call:
interpret(formula = Surv(futime, death) ~ (age + sex + sample.yr +
kappa + lambda)^2, data = flchain, model = rsf_model, na.action = na.pass,
lambda = 0.05)
Model Class: rfsrc, grow, surv
Intercept: 0.99963, 0.99558, 0.99345, ...
Main Effects:
5 main effect terms
Interactions:
10 interaction terms
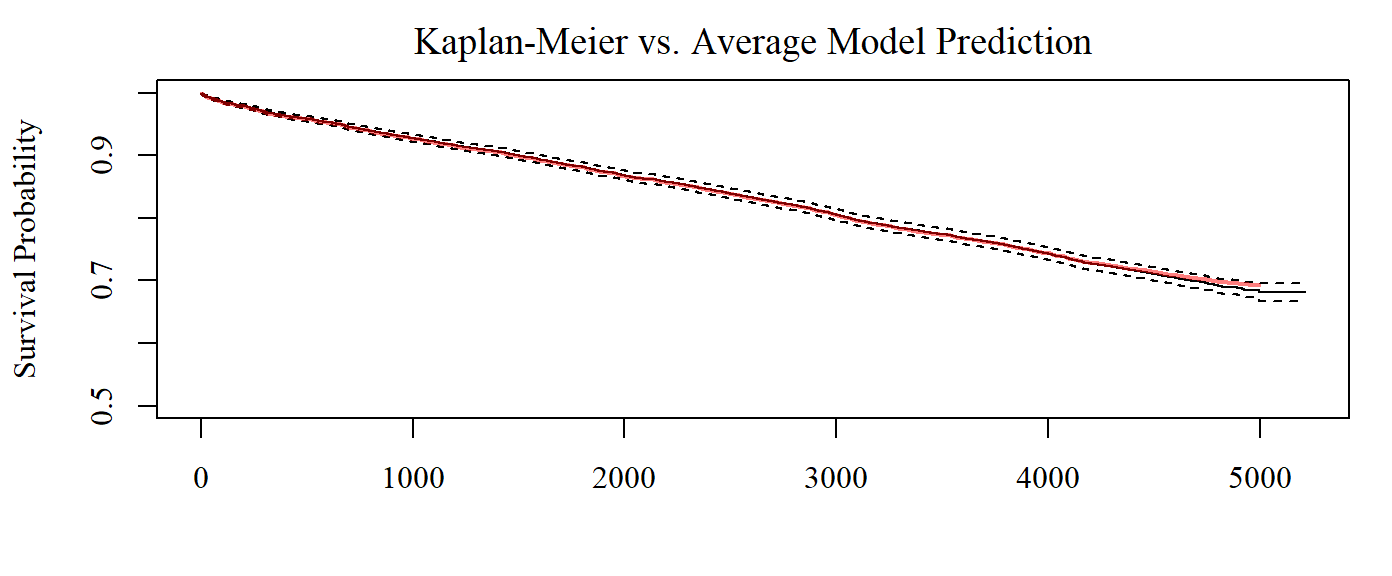
Uninterpreted Variation Ratio: 0.78773, 0.61879, 0.44844, ...Baseline survival probabilities, i.e. the average predicted survival probabilities, are stored as $intercept.
Here, we compare the intercepts with the Kaplan-Meier curve.
km_fit <- survfit(
Surv(futime, death) ~ 1, data = flchain
)
par.midr()
plot(
km_fit,
main = "Kaplan-Meier vs. Average Model Prediction",
# xlab = "Days from Sample Collection",
ylab = "Survival Probability",
ylim = c(0.5, 1.0)
)
lines(
x = as.numeric(labels(mid_model)),
y = mid_model$intercept,
col = "#FF000080",
lwd = 2
)
Since {midr} fits as many models as the number of time periods of interest, the uninterpreted variation ratio changes with time.
Just as survival probabilities change over time, so does the relative impact of each predictor. What strongly influences short-term survival may not be the primary driver of long-term outcomes.
For the MID-based surrogate models, mid.importance() function captures this dynamic behavior, allowing you to visualize how the importance of individual features and their interactions evolves across the specified time horizon.
imp <- mid.importance(mid_model)
p <- ggmid(
imp,
theme = "magma",
terms = rev(head(mid.terms(imp), 10))
) +
theme(legend.position = "none") +
labs(subtitle = "Feature Importance") +
coord_flip()
q <- ggmid(
imp,
type = "series",
theme = "Tableau 10",
terms = mid.terms(imp, interactions = FALSE),
linewidth = 3/4
) +
labs(x = "Days from Sample Collection") +
theme(legend.position = "bottom")
p / q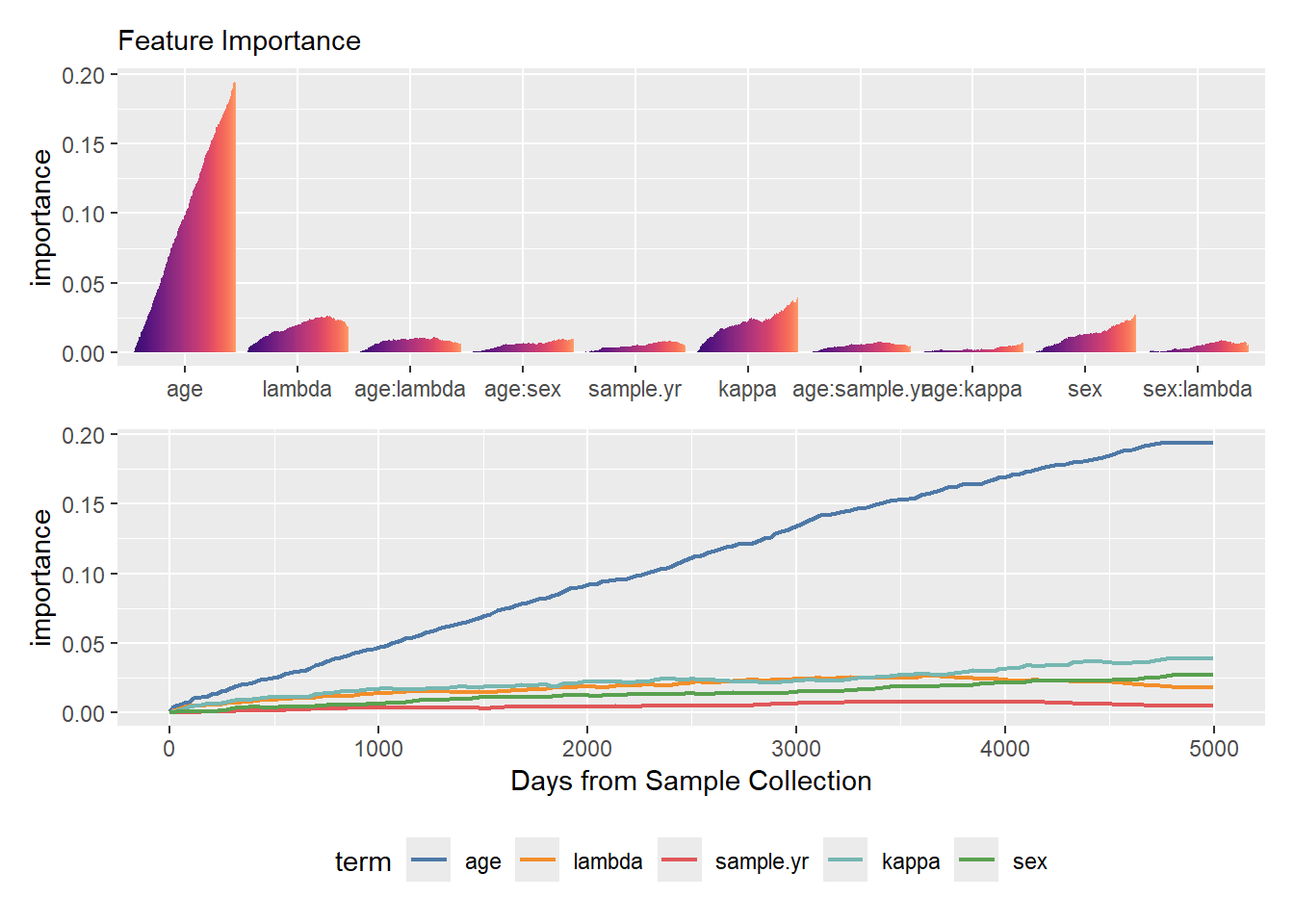
Beyond knowing which features matter, it is crucial to understand how they influence predictions over time. By evaluating the component functions, you can extract the estimated main and interaction effects on survival probabilities. This provides a clear, time-varying view of partial dependencies, showing exactly how different values of a feature shift the survival curve.
plots <- mid.plots(
mid_model,
terms = c("age", "lambda", "kappa", "sex"),
type = "series",
theme = "mako",
resolution = 100,
intercept = TRUE,
limits = c(-0.05, 1.05)
)
plots[[1]] <- plots[[1]] +
labs(subtitle = "Feature Effect on Survival Probability")
plots[[4]] <- plots[[4]] +
labs(x = "Days from Sample Collection")
plots[[1]] / plots[[2]] / plots[[3]] / plots[[4]]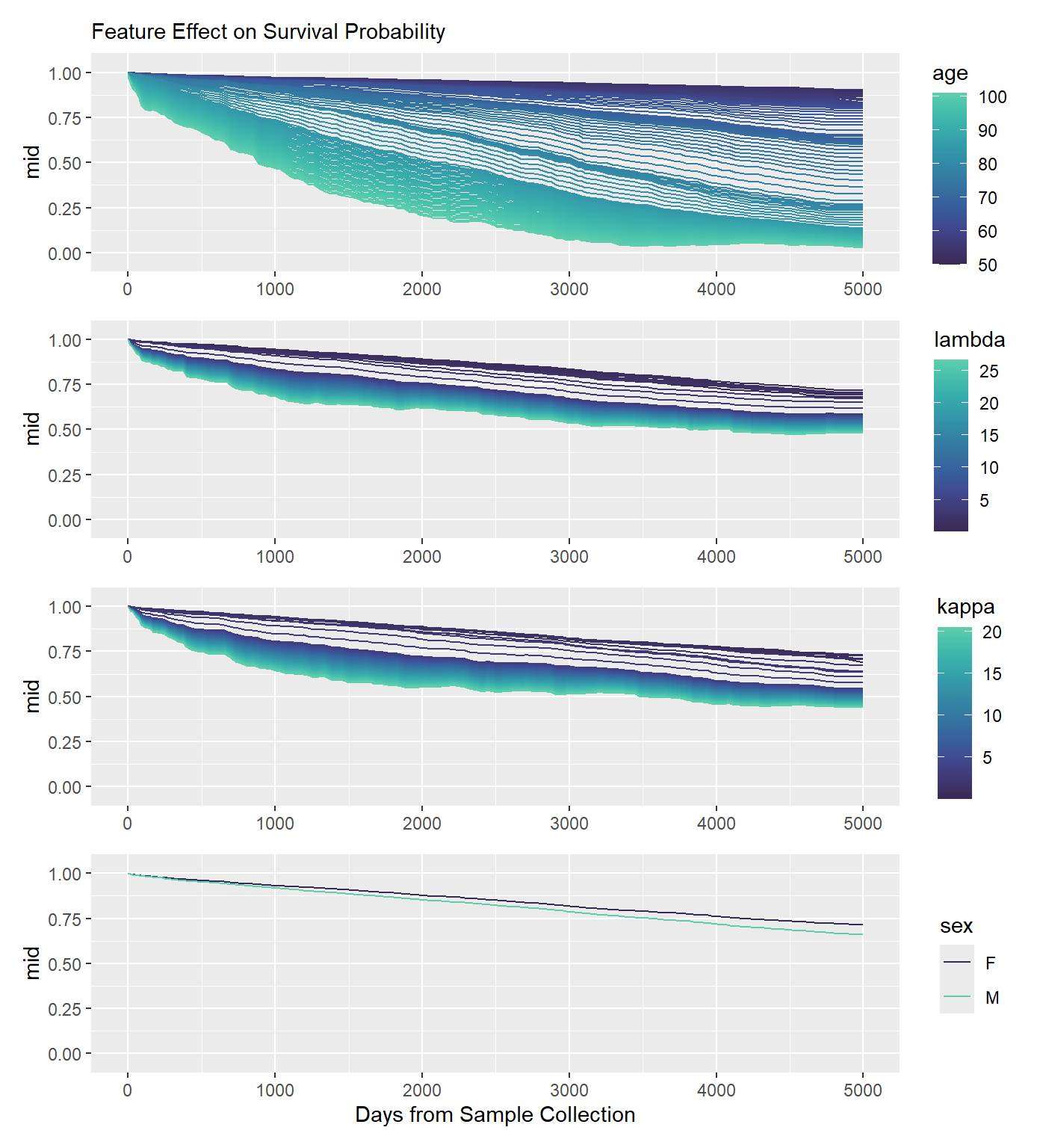
To build trust in clinical or business applications, we often need to explain individual predictions.
The mid.breakdown() function zooms in on a single observation, decomposing that specific patient’s predicted survival curve into additive contributions from the baseline and each feature. This makes it intuitively clear exactly which risk factors are driving an individual’s prognosis at any given moment.
brk <- mid.breakdown(mid_model, data = flchain[1 ,])
p <- ggmid(
brk,
theme = "cividis",
terms = rev(head(mid.terms(imp, interactions = FALSE)))
) +
theme(legend.position = "none") +
labs(subtitle = "Breakdown of Prediction") +
coord_flip()
q <- ggmid(
brk,
type = "series",
theme = "Tableau 10",
terms = mid.terms(imp, interactions = FALSE),
linewidth = 3/4
) +
labs(x = "Days from Sample Collection") +
theme(legend.position = "bottom")
p / q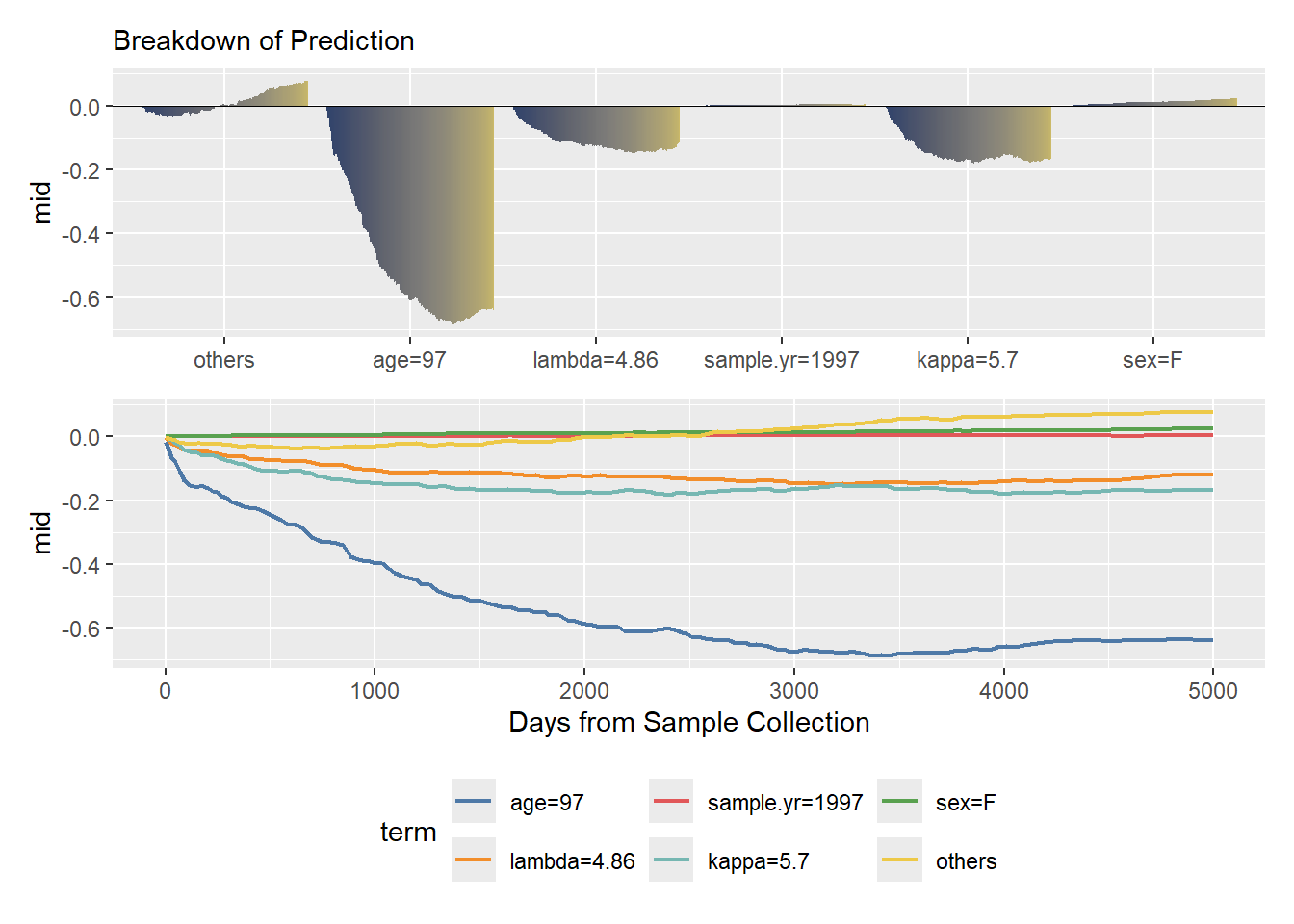
For a more granular look at how a model behaves, Individual Conditional Expectation (ICE) plots are an invaluable tool.
Using mid.conditional(), you can explore counterfactual “what-if” scenarios—visualizing how modifying a specific feature alters the predicted survival outcome for an individual patient. This highlights the change of feature effects in time.