This article presents examples of interpreting survival models using the midr package.
As a fundamental approach to interpreting survival models,
midr first utilizes a prediction function to output a
matrix of survival probabilities,
,
for each observation
at each time point
,
based on the model and the data with the
pred.fun (which defaults to get.yhat()). It
then constructs a series of MID models targeting each column of this
predicted matrix (i.e., the survival probability for each survival time
).
The resulting collection of models across multiple time points is aggregated into an object of class “midrib”. This object is structured to internally share computational resources, such as the design matrix for the calculation of effects. This architecture enables the efficient and smooth extraction of feature effects as they evolve over time.
Cox Proportional Hazard Model
library(survival)
fit_cox <- coxph(
Surv(time, status) ~ .,
data = veteran
)
mid_cox <- interpret(
Surv(time, status) ~ .,
data = veteran,
model = fit_cox,
lambda = .1
)
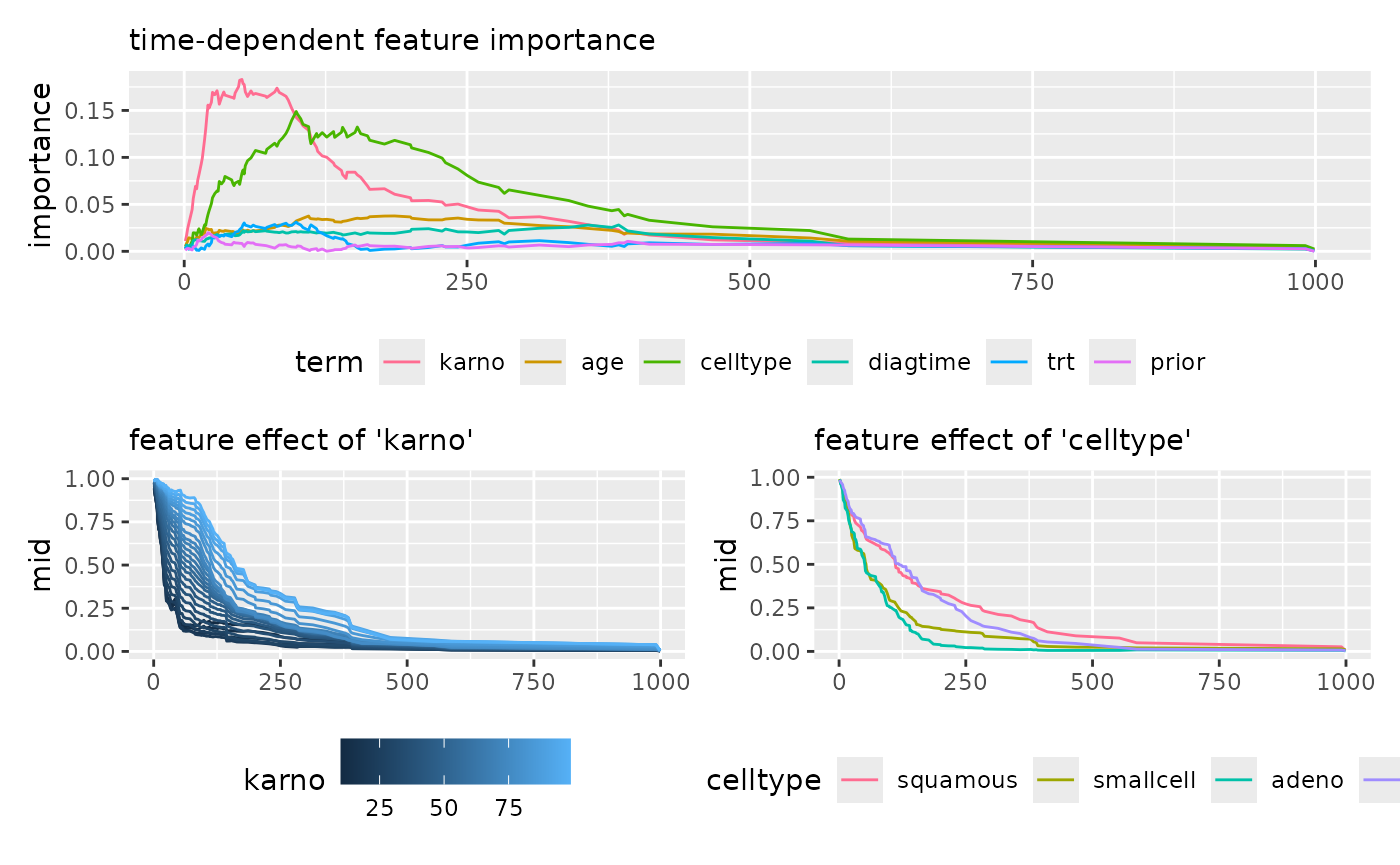
imp_cox <- mid.importance(mid_cox)
p1 <- ggmid(imp_cox, type = "series") +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "time-dependent feature importance")
p2 <- ggmid(mid_cox, term = "karno", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
labs(subtitle = "feature effect of 'karno'")
p3 <- ggmid(mid_cox, term = "celltype", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "feature effect of 'celltype'")
p1 / (p2 + p3)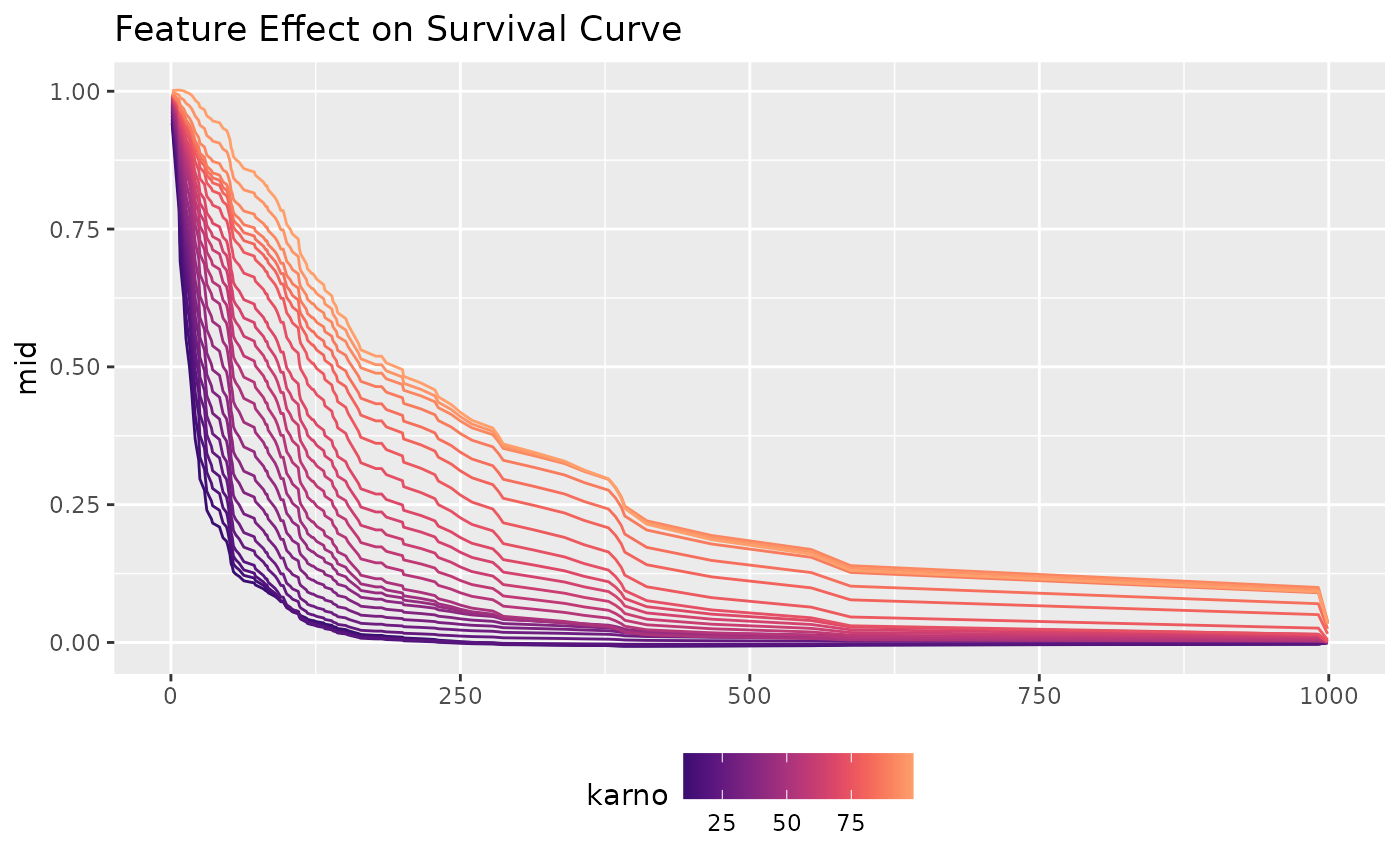
Parametric Survival Model
library(flexsurv)
fit_flex <- flexsurvreg(
Surv(time, status) ~ .,
data = veteran,
dist = "weibull"
)
mid_flex <- interpret(
Surv(time, status) ~ .,
data = veteran,
model = fit_flex,
lambda = .1
)
imp_flex <- mid.importance(mid_flex)
p1 <- ggmid(imp_flex, type = "series") +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "time-dependent feature importance")
p2 <- ggmid(mid_flex, term = "karno", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
labs(subtitle = "feature effect of 'karno'")
p3 <- ggmid(mid_flex, term = "celltype", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "feature effect of 'celltype'")
p1 / (p2 + p3)
Model Based Boosting
library(mboost)
fit_mboost <- glmboost(
Surv(time, status) ~ .,
data = veteran,
family = CoxPH()
)
mid_mboost <- interpret(
Surv(time, status) ~ .,
data = veteran,
model = fit_mboost,
lambda = .1
)
imp_mboost <- mid.importance(mid_mboost)
p1 <- ggmid(imp_mboost, type = "series") +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "time-dependent feature importance")
p2 <- ggmid(mid_mboost, term = "karno", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
labs(subtitle = "feature effect of 'karno'")
p3 <- ggmid(mid_mboost, term = "celltype", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "feature effect of 'celltype'")
p1 / (p2 + p3)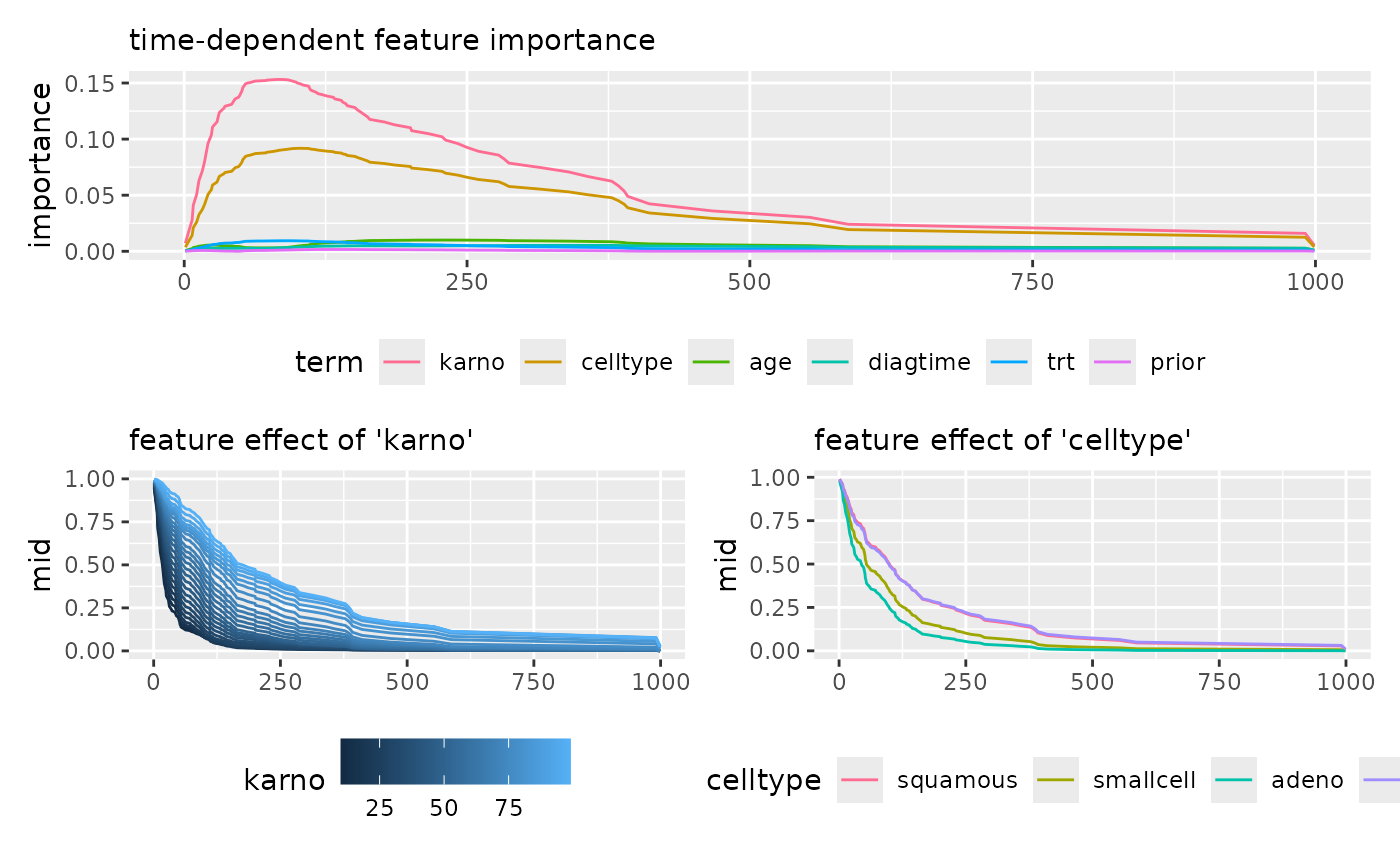
Random Survival Forest
library(randomForestSRC)
fit_rsf <- rfsrc(
Surv(time, status) ~ .,
data = veteran
)
mid_rsf <- interpret(
Surv(time, status) ~ .,
data = veteran,
model = fit_rsf,
lambda = .1
)
imp_rsf <- mid.importance(mid_rsf)
p1 <- ggmid(imp_rsf, type = "series") +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "time-dependent feature importance")
p2 <- ggmid(mid_rsf, term = "karno", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
labs(subtitle = "feature effect of 'karno'")
p3 <- ggmid(mid_rsf, term = "celltype", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "feature effect of 'celltype'")
p1 / (p2 + p3)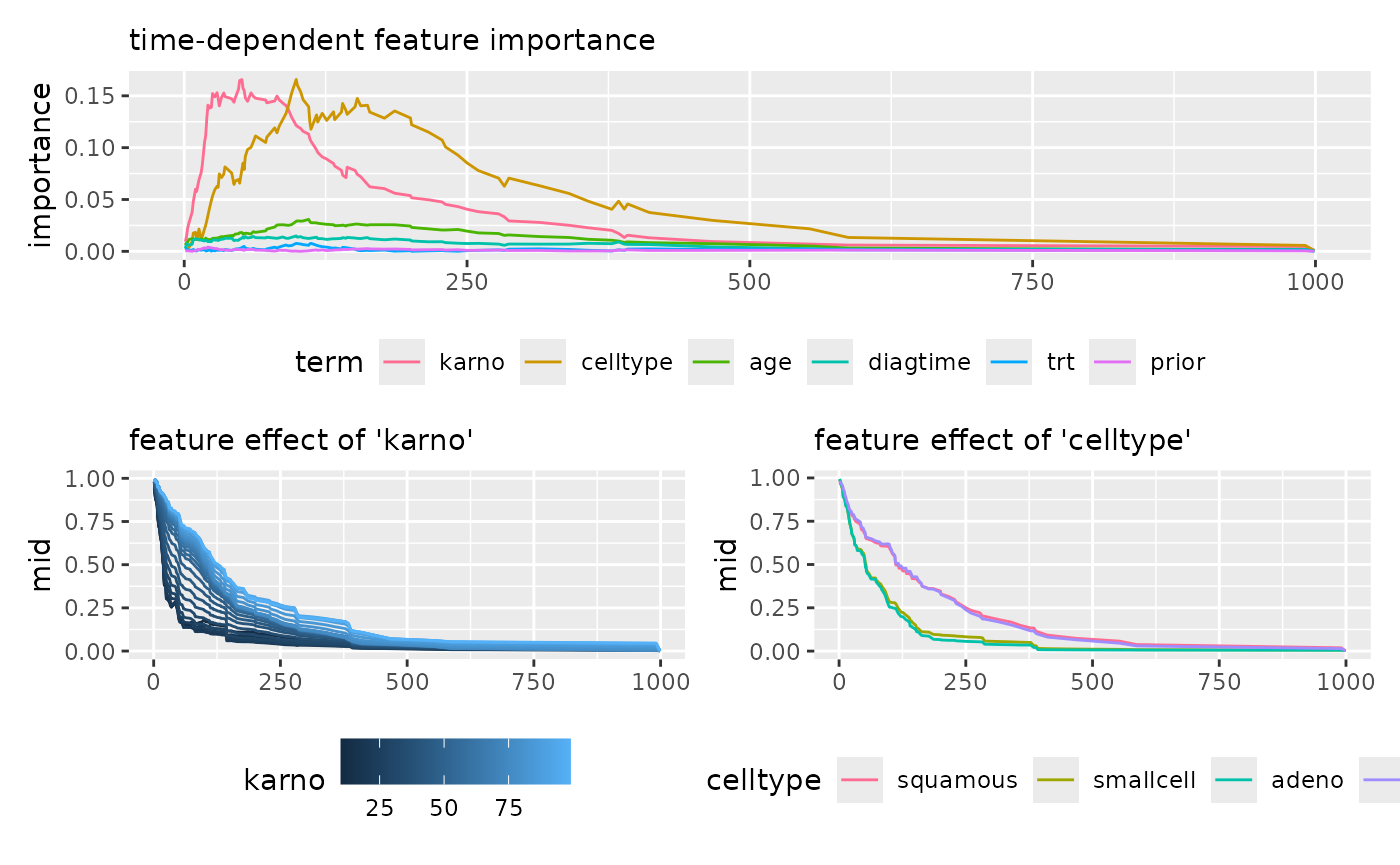
Oblique Random Survival Forest
library(aorsf)
fit_orsf <- orsf(
Surv(time, status) ~ .,
data = veteran
)
mid_orsf <- interpret(
Surv(time, status) ~ .,
data = veteran,
model = fit_orsf,
lambda = .1
)
imp_orsf <- mid.importance(mid_orsf)
p1 <- ggmid(imp_orsf, type = "series") +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "time-dependent feature importance")
p2 <- ggmid(mid_orsf, term = "karno", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
labs(subtitle = "feature effect of 'karno'")
p3 <- ggmid(mid_orsf, term = "celltype", type = "series", intercept = TRUE) +
theme(legend.position = "bottom") +
guides(color = guide_legend(nrow = 1)) +
labs(subtitle = "feature effect of 'celltype'")
p1 / (p2 + p3)